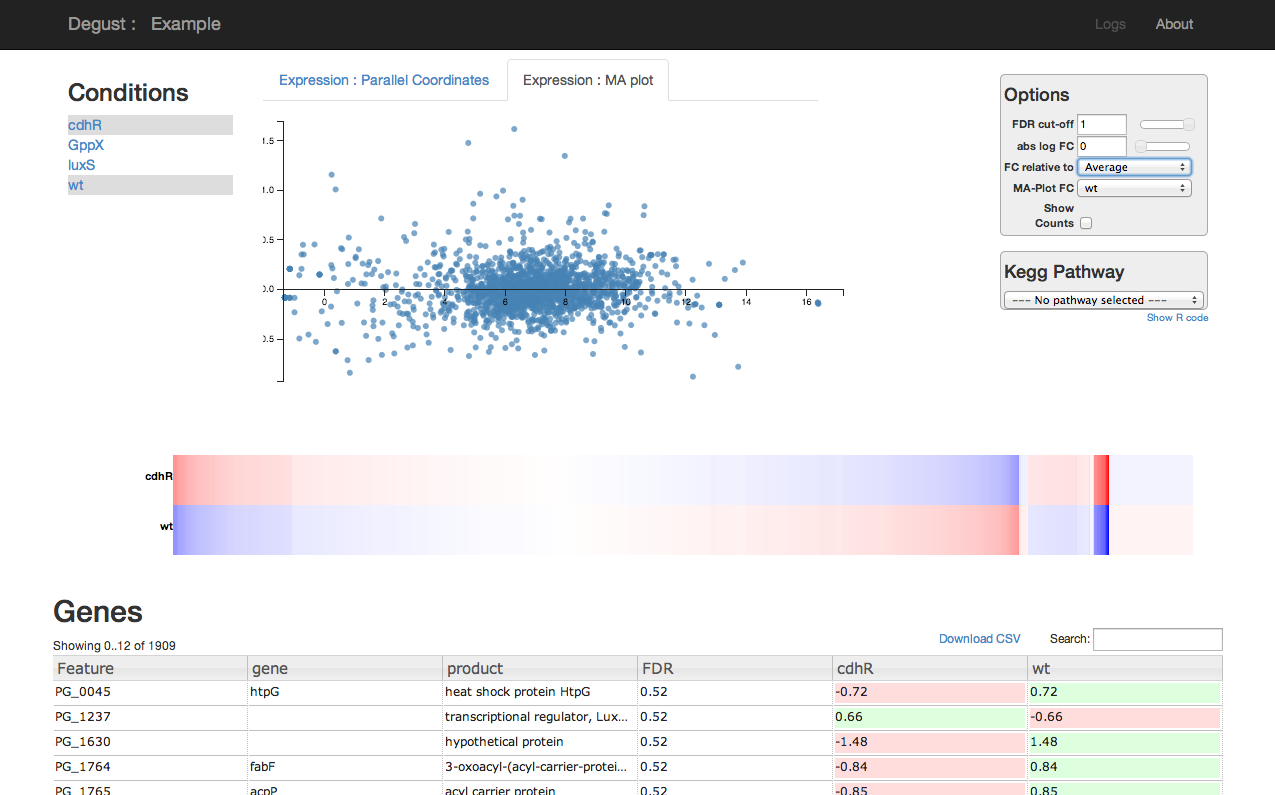
Degust (formerly DGE-Vis)
An interactive web tool for visualising Differential Gene Expression data
or watch a screencast of Degust in action
Using Degust
There are three alternative ways to use Degust. Briefly, these are:
- Upload a CSV of counts per gene to our public server.
- Do your own analysis, and upload a CSV of your analysis to our public server.
- Do your own analysis and create a single HTML file to view and share using degust.py.
Use Degust with per gene read counts
Create your CSV file with read counts per gene - the CSV file requirements are documented on the upload page. You'll then be directed to web page to specify the columns of your CSV including which samples are replicates. The back-end will then analyse your data using voom/limma, and you can visualise the results with Degust.
Using Degust in this manner has the advantage that you can select, within Degust, the conditions you wish to compare. That is, selected 2 samples and a moderated t-test is performed for each gene. Select 3 or more samples and a moderated F-test is used.
Use Degust with pre-analysed data
Perform your own differential expression analysis and produce a CSV file of your results - the CSV file requirements are documented on the upload page. You can then specify which columns are which, and visualise your results with Degust.
Use Degust as a single HTML
Perform your own differential expression analysis and produce a CSV file of your results. You then use a python script, degust.py, to embed your CSV file in an HTML file that can be directly loaded by in browser.
Example using Degust from a LIMMA analysis
Assuming you have analysed your data using R with voom/limma, and have a MArrayLM object containing your results you can use Degust as follows.
> # Save our analysis object 'efit' to a CSV file
> class(efit)
[1] "MArrayLM"
attr(,"package")
[1] "limma"
> colnames(efit)
[1] "GppX" "luxS" "cdhR"
> write.csv(topTable(efit, number=Inf), 'dge.csv', row.names=F)
> quit()
bash$
The file we created above, dge.csv, could be loaded directly onto the public server.
Alternatively, you can create your own HTML file using degust.py as follows:
bash$ python degust.py -o dge.html --name 'My DGE experiment' \
--primary 'WT' --avg AveExpr --fdr adj.P.Val \
--logFC "GppX,luxS,cdhR" \
--info "Feature,product" dge.csv
Contact
Degust was written by David R. Powell (@d_r_powell).
Please raise any issues on the Github issues page
Degust is supported in part by